Ethyl-ferulate
General
Type : Acrylate || Feruloyl || Hydroxycinnamic-ester
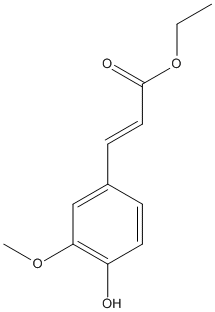
Chemical_Nomenclature : ethyl (E)-3-(4-hydroxy-3-methoxyphenyl)prop-2-enoate
Canonical SMILES : CCOC(=O)C=CC1=CC(=C(C=C1)O)OC
InChI : InChI=1S\/C12H14O4\/c1-3-16-12(14)7-5-9-4-6-10(13)11(8-9)15-2\/h4-8,13H,3H2,1-2H3\/b7-5+
InChIKey : ATJVZXXHKSYELS-FNORWQNLSA-N
Other name(s) : Ethyl ferulate, Ferulic acid ethyl ester, Ethyl 3-(4-hydroxy-3-methoxyphenyl)acrylate, Ethyl 4'-hydroxy-3'-methoxycinnamate, UNII-5B8915UELW
Target
References (8)
| Title : Feruloyl Esterase (LaFae) from Lactobacillus acidophilus: Structural Insights and Functional Characterization for Application in Ferulic Acid Production - Jeon_2023_Int.J.Mol.Sci_24_11170 |
| Author(s) : Jeon S , Hwang J , Do H , Le L , Lee CW , Yoo W , Lee MJ , Shin SC , Kim KK , Kim HW , Lee JH |
| Ref : Int J Mol Sci , 24 :11170 , 2023 |
| Abstract : Jeon_2023_Int.J.Mol.Sci_24_11170 |
| ESTHER : Jeon_2023_Int.J.Mol.Sci_24_11170 |
| PubMedSearch : Jeon_2023_Int.J.Mol.Sci_24_11170 |
| PubMedID: 37446348 |
| Gene_locus related to this paper: lacac-q5fi30 |
| Title : Structure-guided rational design of the Geobacillus thermoglucosidasius feruloyl esterase GthFAE to improve its thermostability - Yang_2022_Biochem.Biophys.Res.Commun_600_117 |
| Author(s) : Yang W , Sun L , Dong P , Chen Y , Zhang H , Huang X , Wu L , Chen L , Jing D , Wu Y |
| Ref : Biochemical & Biophysical Research Communications , 600 :117 , 2022 |
| Abstract : Yang_2022_Biochem.Biophys.Res.Commun_600_117 |
| ESTHER : Yang_2022_Biochem.Biophys.Res.Commun_600_117 |
| PubMedSearch : Yang_2022_Biochem.Biophys.Res.Commun_600_117 |
| PubMedID: 35219099 |
| Gene_locus related to this paper: partm-GthFAE |
| Title : Loop of Streptomyces Feruloyl Esterase Plays an Important Role in the Enzyme's Catalyzing the Release of Ferulic Acid from Biomass - Uraji_2018_Appl.Environ.Microbiol_84_ |
| Author(s) : Uraji M , Tamura H , Mizohata E , Arima J , Wan K , Ogawa K , Inoue T , Hatanaka T |
| Ref : Applied Environmental Microbiology , 84 : , 2018 |
| Abstract : Uraji_2018_Appl.Environ.Microbiol_84_ |
| ESTHER : Uraji_2018_Appl.Environ.Microbiol_84_ |
| PubMedSearch : Uraji_2018_Appl.Environ.Microbiol_84_ |
| PubMedID: 29150515 |
| Gene_locus related to this paper: strcj-estA |
| Title : Characterization of a feruloyl esterase from Aspergillus terreus facilitates the division of fungal enzymes from Carbohydrate Esterase family 1 of the carbohydrate-active enzymes (CAZy) database - Makela_2018_Microb.Biotechnol_11_869 |
| Author(s) : Makela MR , Dilokpimol A , Koskela SM , Kuuskeri J , de Vries RP , Hilden K |
| Ref : Microb Biotechnol , 11 :869 , 2018 |
| Abstract : Makela_2018_Microb.Biotechnol_11_869 |
| ESTHER : Makela_2018_Microb.Biotechnol_11_869 |
| PubMedSearch : Makela_2018_Microb.Biotechnol_11_869 |
| PubMedID: 29697197 |
| Gene_locus related to this paper: asptn-q0ci40 |
| Title : Characterization of Cinnamoyl Esterases from Different Lactobacilli and Bifidobacteria - Fritsch_2017_Curr.Microbiol_74_247 |
| Author(s) : Fritsch C , Jansch A , Ehrmann MA , Toelstede S , Vogel RF |
| Ref : Curr Microbiol , 74 :247 , 2017 |
| Abstract : Fritsch_2017_Curr.Microbiol_74_247 |
| ESTHER : Fritsch_2017_Curr.Microbiol_74_247 |
| PubMedSearch : Fritsch_2017_Curr.Microbiol_74_247 |
| PubMedID: 27999938 |
| Gene_locus related to this paper: limf3-b2gdc2 , lache-u6f2k7 , bifa0-b8du06 , lacac-q5fi30 , lacga-q040s2 , lacpl-LP.2953 , lacre-q2bvr2 |
| Title : An inserted alpha\/beta subdomain shapes the catalytic pocket of Lactobacillus johnsonii cinnamoyl esterase - Lai_2011_PLoS.One_6_e23269 |
| Author(s) : Lai KK , Stogios PJ , Vu C , Xu X , Cui H , Molloy S , Savchenko A , Yakunin A , Gonzalez CF |
| Ref : PLoS ONE , 6 :e23269 , 2011 |
| Abstract : Lai_2011_PLoS.One_6_e23269 |
| ESTHER : Lai_2011_PLoS.One_6_e23269 |
| PubMedSearch : Lai_2011_PLoS.One_6_e23269 |
| PubMedID: 21876742 |
| Gene_locus related to this paper: lacjo-q74hk5 |
| Title : Structural and functional characterization of a promiscuous feruloyl esterase (Est1E) from the rumen bacterium Butyrivibrio proteoclasticus - Goldstone_2010_Proteins_78_1457 |
| Author(s) : Goldstone DC , Villas-Boas SG , Till M , Kelly WJ , Attwood GT , Arcus VL |
| Ref : Proteins , 78 :1457 , 2010 |
| Abstract : Goldstone_2010_Proteins_78_1457 |
| ESTHER : Goldstone_2010_Proteins_78_1457 |
| PubMedSearch : Goldstone_2010_Proteins_78_1457 |
| PubMedID: 20058325 |
| Gene_locus related to this paper: 9firm-Est1E |
| Title : Identification of a type-D feruloyl esterase from Neurospora crassa - Crepin_2004_Appl.Microbiol.Biotechnol_63_567 |
| Author(s) : Crepin VF , Faulds CB , Connerton IF |
| Ref : Applied Microbiology & Biotechnology , 63 :567 , 2004 |
| Abstract : Crepin_2004_Appl.Microbiol.Biotechnol_63_567 |
| ESTHER : Crepin_2004_Appl.Microbiol.Biotechnol_63_567 |
| PubMedSearch : Crepin_2004_Appl.Microbiol.Biotechnol_63_567 |
| PubMedID: 14595525 |
| Gene_locus related to this paper: neucr-FAED |