Ac-Leu-p-nitroanilide
General
Type : Peptide || p-nitroaniline || pNP || Chromogen
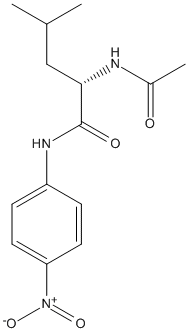
Chemical_Nomenclature : (2S)-2-acetamido-4-methyl-N-(4-nitrophenyl)pentanamide
Canonical SMILES : CC(C)CC(C(=O)NC1=CC=C(C=C1)[N+](=O)[O-])NC(=O)C
InChI : InChI=1S\/C14H19N3O4\/c1-9(2)8-13(15-10(3)18)14(19)16-11-4-6-12(7-5-11)17(20)21\/h4-7,9,13H,8H2,1-3H3,(H,15,18)(H,16,19)\/t13-\/m0\/s1
InChIKey : SLJKQHDBYBGOMH-ZDUSSCGKSA-N
Other name(s) : Ac-Leu-pNA, N-Acetyl-L-leucine-p-nitroanilide, Acetyl-L-leucine 4-nitroanilide, AC1OLQVE, CTK8F7540
Target
Families : ACPH_Peptidase_S9
References (3)
| Title : Understanding the interactions of different substrates with wild-type and mutant acylaminoacyl peptidase using molecular dynamics simulations - Zhu_2017_J.Biomol.Struct.Dyn__1 |
| Author(s) : Zhu J , Wang Y , Li X , Han W , Zhao L |
| Ref : J Biomol Struct Dyn , :1 , 2017 |
| Abstract : Zhu_2017_J.Biomol.Struct.Dyn__1 |
| ESTHER : Zhu_2017_J.Biomol.Struct.Dyn__1 |
| PubMedSearch : Zhu_2017_J.Biomol.Struct.Dyn__1 |
| PubMedID: 29235404 |
| Gene_locus related to this paper: aerpe-APE1547 |
| Title : Exploration of the chlorpyrifos escape pathway from acylpeptide hydrolases using steered molecular dynamics simulations - Wang_2016_J.Biomol.Struct.Dyn_34_749 |
| Author(s) : Wang D , Jin H , Wang J , Guan S , Zhang Z , Han W |
| Ref : J Biomol Struct Dyn , 34 :749 , 2016 |
| Abstract : Wang_2016_J.Biomol.Struct.Dyn_34_749 |
| ESTHER : Wang_2016_J.Biomol.Struct.Dyn_34_749 |
| PubMedSearch : Wang_2016_J.Biomol.Struct.Dyn_34_749 |
| PubMedID: 26155973 |
| Gene_locus related to this paper: aerpe-APE1547 |
| Title : Discrimination of esterase and peptidase activities of acylaminoacyl peptidase from hyperthermophilic Aeropyrum pernix K1 by a single mutation - Wang_2006_J.Biol.Chem_281_18618 |
| Author(s) : Wang Q , Yang G , Liu Y , Feng Y |
| Ref : Journal of Biological Chemistry , 281 :18618 , 2006 |
| Abstract : Wang_2006_J.Biol.Chem_281_18618 |
| ESTHER : Wang_2006_J.Biol.Chem_281_18618 |
| PubMedSearch : Wang_2006_J.Biol.Chem_281_18618 |
| PubMedID: 16670095 |
| Gene_locus related to this paper: aerpe-APE1547 |