AHMMCE
General
Type : Cannabinoid-Receptor-ligand
Chemical_Nomenclature :
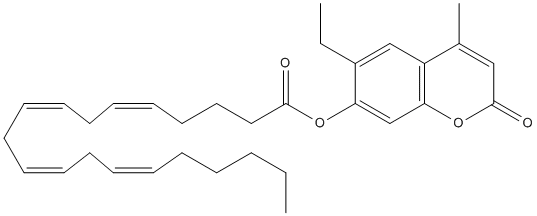
Canonical SMILES : C(C=CCC=CCCCCC)C=CCC=CCCCC(=O)OC1=CC2=C(C=C1CC)C(=CC(O2)=O)C
InChI : InChI=1S\/C32H42O4\/c1-4-6-7-8-9-10-11-12-13-14-15-16-17-18-19-20-21-22-31(33)35-29-25-30-28(24-27(29)5-2)26(3)23-32(34)36-30\/h9-10,12-13,15-16,18-19,23-25H,4-8,11,14,17,20-22H2,1-3H3
InChIKey : PDZLOIANBXDWRG-UHFFFAOYSA-N
Other name(s) :
MW : 490.67
Formula : C32H42O4
CAS_number :
PubChem :
UniChem :
Iuphar :
Target
Families : Monoglyceridelipase_lysophospholip, ABHD6-Lip
References (5)
| Title : Biochemical and Proteomic Characterization of Recombinant Human alpha\/beta Hydrolase Domain 6 - Shields_2019_Sci.Rep_9_890 |
| Author(s) : Shields CM , Zvonok N , Zvonok A , Makriyannis A |
| Ref : Sci Rep , 9 :890 , 2019 |
| Abstract : Shields_2019_Sci.Rep_9_890 |
| ESTHER : Shields_2019_Sci.Rep_9_890 |
| PubMedSearch : Shields_2019_Sci.Rep_9_890 |
| PubMedID: 30696836 |
| Gene_locus related to this paper: human-ABHD6 |
| Title : The role of human monoacylglycerol lipase (hMAGL) binding pocket in breakup of unsaturated phospholipid membranes - Karageorgos_2017_Anal.Biochem_536_90 |
| Author(s) : Karageorgos I , Silin VI , Zvonok N , Marino J , Janero DR , Makriyannis A |
| Ref : Analytical Biochemistry , 536 :90 , 2017 |
| Abstract : Karageorgos_2017_Anal.Biochem_536_90 |
| ESTHER : Karageorgos_2017_Anal.Biochem_536_90 |
| PubMedSearch : Karageorgos_2017_Anal.Biochem_536_90 |
| PubMedID: 28822686 |
| Gene_locus related to this paper: human-MGLL |
| Title : Active-site inhibitors modulate the dynamic properties of human monoacylglycerol lipase: a hydrogen exchange mass spectrometry study - Karageorgos_2013_Biochemistry_52_5016 |
| Author(s) : Karageorgos I , Wales TE , Janero DR , Zvonok N , Vemuri VK , Engen JR , Makriyannis A |
| Ref : Biochemistry , 52 :5016 , 2013 |
| Abstract : Karageorgos_2013_Biochemistry_52_5016 |
| ESTHER : Karageorgos_2013_Biochemistry_52_5016 |
| PubMedSearch : Karageorgos_2013_Biochemistry_52_5016 |
| PubMedID: 23795559 |
| Title : Membrane phospholipid bilayer as a determinant of monoacylglycerol lipase kinetic profile and conformational repertoire - Nasr_2013_Protein.Sci_22_774 |
| Author(s) : Nasr ML , Shi X , Bowman AL , Johnson M , Zvonok N , Janero DR , Vemuri VK , Wales TE , Engen JR , Makriyannis A |
| Ref : Protein Science , 22 :774 , 2013 |
| Abstract : Nasr_2013_Protein.Sci_22_774 |
| ESTHER : Nasr_2013_Protein.Sci_22_774 |
| PubMedSearch : Nasr_2013_Protein.Sci_22_774 |
| PubMedID: 23553709 |
| Title : Endocannabinoid enzyme engineering: soluble human thio-monoacylglycerol lipase (sol-S-hMGL) - Karageorgos_2012_ACS.Chem.Neurosci_3_393 |
| Author(s) : Karageorgos I , Zvonok N , Janero DR , Vemuri VK , Shukla V , Wales TE , Engen JR , Makriyannis A |
| Ref : ACS Chem Neurosci , 3 :393 , 2012 |
| Abstract : Karageorgos_2012_ACS.Chem.Neurosci_3_393 |
| ESTHER : Karageorgos_2012_ACS.Chem.Neurosci_3_393 |
| PubMedSearch : Karageorgos_2012_ACS.Chem.Neurosci_3_393 |
| PubMedID: 22860208 |
| Gene_locus related to this paper: human-MGLL |