5MXP
Name : Haloalkane dehalogenase DmxA from Marinobacter sp. ELB17 possessing a unique catalytic residue
Revelation date : 25-Jul-2018
Family : Haloalkane_dehalogenase-HLD1
Gene_locus : 9alte-a3jb27
PDB file : ESTHER: header of PDB entry RCSB: Full entry
Comment
Chrast, L., Tratsiak, K., Daniel, L., Prudnikova, T., Brezovsky, J., Smatanova, I.K., Chaloupkova, R., Damborsky, J.Structural basis of paradoxically high thermostability of dehalogenase from psychrophilic bacterium
Ligand :
References (2)
| Title : Deciphering the Structural Basis of High Thermostability of Dehalogenase from Psychrophilic Bacterium Marinobacter sp. ELB17 - Chrast_2019_Microorganisms_7_ |
| Author(s) : Chrast L , Tratsiak K , Planas-Iglesias J , Daniel L , Prudnikova T , Brezovsky J , Bednar D , Kuta Smatanova I , Chaloupkova R , Damborsky J |
| Ref : Microorganisms , 7 : , 2019 |
| Abstract : Chrast_2019_Microorganisms_7_ |
| ESTHER : Chrast_2019_Microorganisms_7_ |
| PubMedSearch : Chrast_2019_Microorganisms_7_ |
| PubMedID: 31661858 |
| Gene_locus related to this paper: 9alte-a3jb27 |
| Title : Crystallographic analysis of new psychrophilic haloalkane dehalogenases: DpcA from Psychrobacter cryohalolentis K5 and DmxA from Marinobacter sp. ELB17 - Tratsiak_2013_Acta.Crystallogr.Sect.F.Struct.Biol.Cryst.Commun_69_683 |
| Author(s) : Tratsiak K , Degtjarik O , Drienovska I , Chrast L , Rezacova P , Kuty M , Chaloupkova R , Damborsky J , Kuta Smatanova I |
| Ref : Acta Crystallographica Sect F Struct Biol Cryst Commun , 69 :683 , 2013 |
| Abstract : Tratsiak_2013_Acta.Crystallogr.Sect.F.Struct.Biol.Cryst.Commun_69_683 |
| ESTHER : Tratsiak_2013_Acta.Crystallogr.Sect.F.Struct.Biol.Cryst.Commun_69_683 |
| PubMedSearch : Tratsiak_2013_Acta.Crystallogr.Sect.F.Struct.Biol.Cryst.Commun_69_683 |
| PubMedID: 23722854 |
| Gene_locus related to this paper: 9alte-a3jb27 , psyck-q1qbb9 |
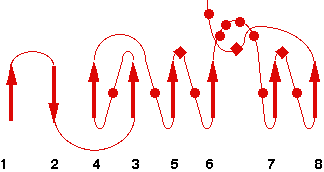
Representative scheme of Prolylcarboxypeptidase structure and an image from PDBsum server