1AQL
Name : Bovine bile-salt-activated lipase + taurocholate
Revelation date : 05-Aug-1998
Family : Cholesterol_esterase
Gene_locus : bovin-balip
PDB file : ESTHER: header of PDB entry RCSB: Full entry
Comment
Ligand :
References (1)
| Title : The crystal structure of bovine bile salt activated lipase: insights into the bile salt activation mechanism - Wang_1997_Structure_5_1209 |
| Author(s) : Wang X , Wang CS , Tang J , Dyda F , Zhang XC |
| Ref : Structure , 5 :1209 , 1997 |
| Abstract : Wang_1997_Structure_5_1209 |
| ESTHER : Wang_1997_Structure_5_1209 |
| PubMedSearch : Wang_1997_Structure_5_1209 |
| PubMedID: 9331420 |
| Gene_locus related to this paper: bovin-balip |
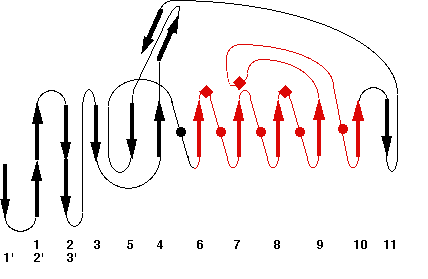
Representative scheme of Prolylcarboxypeptidase structure and an image from PDBsum server