dMUP
Substrate for testing OP-hydrolase activity
General
Type : Organophosphate,Coumarin
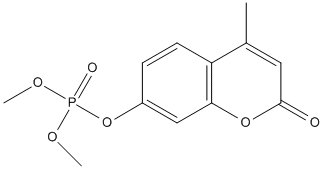
Chemical_Nomenclature : dimethyl (4-methyl-2-oxochromen-7-yl) phosphate
Canonical SMILES : CC1=CC(=O)OC2=C1C=CC(=C2)OP(=O)(OC)OC
InChI : InChI=1S\/C12H13O6P\/c1-8-6-12(13)17-11-7-9(4-5-10(8)11)18-19(14,15-2)16-3\/h4-7H,1-3H3
InChIKey : SVVSCEWVQSATRC-UHFFFAOYSA-N
Other name(s) : dimethyl 4-methylumbelliferyl phosphate,O,O-dimehtyl 4-methylumbelliferyl phosphate,SCHEMBL1023154
MW : 284.20
Formula : C12H13O6P
CAS_number :
PubChem : 60108029
UniChem : SVVSCEWVQSATRC-UHFFFAOYSA-N
IUPHAR :
Wikipedia :
Target
References (2)
| Title : Hydrolysis of organophosphorus insecticides by in vitro modified carboxylesterase E3 from Lucilia cuprina - Heidari_2004_Insect.Biochem.Mol.Biol_34_353 |
| Author(s) : Heidari R , Devonshire AL , Campbell BE , Bell KL , Dorrian SJ , Oakeshott JG , Russell RJ |
| Ref : Insect Biochemistry & Molecular Biology , 34 :353 , 2004 |
| Abstract : Heidari_2004_Insect.Biochem.Mol.Biol_34_353 |
| ESTHER : Heidari_2004_Insect.Biochem.Mol.Biol_34_353 |
| PubMedSearch : Heidari_2004_Insect.Biochem.Mol.Biol_34_353 |
| PubMedID: 15041019 |
| Gene_locus related to this paper: luccu-E3aest7 |
| Title : Kinetic efficiency of mutant carboxylesterases implicated in organophosphate insecticide resistance - Devonshire_2003_Pestic.Biochem.Physiol_76_1 |
| Author(s) : Devonshire AL , Heidari R , Bell KL , Campbell PM , Campbell BE , Odgers WA , Oakeshott JG , Russell RJ |
| Ref : Pesticide Biochemistry and Physiology , 76 :1 , 2003 |
| Abstract : Devonshire_2003_Pestic.Biochem.Physiol_76_1 |
| ESTHER : Devonshire_2003_Pestic.Biochem.Physiol_76_1 |
| PubMedSearch : Devonshire_2003_Pestic.Biochem.Physiol_76_1 |
| PubMedID: |
| Gene_locus related to this paper: luccu-E3aest7 , musdo-EST23aes07 |