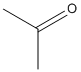
Acetone
General
Type : Ketone
Chemical_Nomenclature : propan-2-one
Canonical SMILES : CC(=O)C
InChI : InChI=1S\/C3H6O\/c1-3(2)4\/h1-2H3
InChIKey : CSCPPACGZOOCGX-UHFFFAOYSA-N
Other name(s) : 2-propanone,Propanone,Dimethyl ketone,Methyl ketone,Dimethylformaldehyde
MW : 58.07
Formula : C3H6O
CAS_number : 67-64-1
PubChem : 180
UniChem : CSCPPACGZOOCGX-UHFFFAOYSA-N
IUPHAR :
Wikipedia :
Target
Families : Acetone ligand of proteins in family: Hydroxynitrile_lyase
Stucture :
Protein : hevbr-hnl || manes-hnl
References (4)
| Title : Atomic resolution crystal structures and quantum chemistry meet to reveal subtleties of hydroxynitrile lyase catalysis - Schmidt_2008_J.Biol.Chem_283_21827 |
| Author(s) : Schmidt A , Gruber K , Kratky C , Lamzin VS |
| Ref : Journal of Biological Chemistry , 283 :21827 , 2008 |
| Abstract : Schmidt_2008_J.Biol.Chem_283_21827 |
| ESTHER : Schmidt_2008_J.Biol.Chem_283_21827 |
| PubMedSearch : Schmidt_2008_J.Biol.Chem_283_21827 |
| PubMedID: 18524775 |
| Gene_locus related to this paper: hevbr-hnl |
| Title : Reaction mechanism of hydroxynitrile lyases of the alpha\/beta-hydrolase superfamily: the three-dimensional structure of the transient enzyme-substrate complex certifies the crucial role of LYS236 - Gruber_2004_J.Biol.Chem_279_20501 |
| Author(s) : Gruber K , Gartler G , Krammer B , Schwab H , Kratky C |
| Ref : Journal of Biological Chemistry , 279 :20501 , 2004 |
| Abstract : Gruber_2004_J.Biol.Chem_279_20501 |
| ESTHER : Gruber_2004_J.Biol.Chem_279_20501 |
| PubMedSearch : Gruber_2004_J.Biol.Chem_279_20501 |
| PubMedID: 14998991 |
| Gene_locus related to this paper: hevbr-hnl |
| Title : Structure of hydroxynitrile lyase from Manihot esculenta in complex with substrates acetone and chloroacetone: implications for the mechanism of cyanogenesis - Lauble_2001_Acta.Crystallogr.D.Biol.Crystallogr_57_194 |
| Author(s) : Lauble H , Forster S , Miehlich B , Wajant H , Effenberger F |
| Ref : Acta Crystallographica D Biol Crystallogr , 57 :194 , 2001 |
| Abstract : Lauble_2001_Acta.Crystallogr.D.Biol.Crystallogr_57_194 |
| ESTHER : Lauble_2001_Acta.Crystallogr.D.Biol.Crystallogr_57_194 |
| PubMedSearch : Lauble_2001_Acta.Crystallogr.D.Biol.Crystallogr_57_194 |
| PubMedID: 11173464 |
| Gene_locus related to this paper: manes-hnl |
| Title : Three-dimensional structures of enzyme-substrate complexes of the hydroxynitrile lyase from Hevea brasiliensis - Zuegg_1999_Protein.Sci_8_1990 |
| Author(s) : Zuegg J , Gruber K , Gugganig M , Wagner UG , Kratky C |
| Ref : Protein Science , 8 :1990 , 1999 |
| Abstract : Zuegg_1999_Protein.Sci_8_1990 |
| ESTHER : Zuegg_1999_Protein.Sci_8_1990 |
| PubMedSearch : Zuegg_1999_Protein.Sci_8_1990 |
| PubMedID: 10548044 |
| Gene_locus related to this paper: hevbr-hnl |