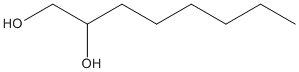
1,2-Octanediol
General
Type :
Chemical_Nomenclature : (2R)-octane-1,2-diol
Canonical SMILES : CCCCCCC(CO)O
InChI : InChI=1S\/C8H18O2\/c1-2-3-4-5-6-8(10)7-9\/h8-10H,2-7H2,1H3\/t8-\/m1\/s1
InChIKey : AEIJTFQOBWATKX-MRVPVSSYSA-N || AEIJTFQOBWATKX-QMMMGPOBSA-N || AEIJTFQOBWATKX-UHFFFAOYSA-N
Other name(s) : (2R)-octane-1,2-diol,(R)-(+)-1,2-Octanediol,(R)-1,2-octanediol,7F2,AC1ODVSI,(2S)-octane-1,2-diol,(S)-1,2-OCTANEDIOL,(S)-(-)-1,2-Octanediol,87720-91-0,J-640384,AC1OE5RZ,1,2-OCTANEDIOL,1117-86-8,Octane-1,2-diol,1,2-Dihydroxyoctane,1,2-Octylene glycol,Capyryl Glycol
MW : 146.23
Formula : C8H18O2
CAS_number : 87720-90-9
PubChem : 6994435, 6999933, 14231
UniChem : AEIJTFQOBWATKX-MRVPVSSYSA-N || AEIJTFQOBWATKX-QMMMGPOBSA-N || AEIJTFQOBWATKX-UHFFFAOYSA-N
IUPHAR :
Wikipedia :
Target
Families : 1,2-Octanediol ligand of proteins in family: CFTR-inhibitory-factor_Cif
Stucture :
Protein : pseae-PA2934
References (2)
| Title : Potential of Y. lipolytica epoxide hydrolase for efficient production of enantiopure (R)-1,2-octanediol - Godase_2023_AMB.Express_13_77 |
| Author(s) : Godase VP , Kumar VR , Kumar AR |
| Ref : AMB Express , 13 :77 , 2023 |
| Abstract : Godase_2023_AMB.Express_13_77 |
| ESTHER : Godase_2023_AMB.Express_13_77 |
| PubMedSearch : Godase_2023_AMB.Express_13_77 |
| PubMedID: 37495892 |
| Title : Active-Site Flexibility and Substrate Specificity in a Bacterial Virulence Factor: Crystallographic Snapshots of an Epoxide Hydrolase - Hvorecny_2017_Structure_25_697 |
| Author(s) : Hvorecny KL , Bahl CD , Kitamura S , Lee KSS , Hammock BD , Morisseau C , Madden DR |
| Ref : Structure , 25 :697 , 2017 |
| Abstract : Hvorecny_2017_Structure_25_697 |
| ESTHER : Hvorecny_2017_Structure_25_697 |
| PubMedSearch : Hvorecny_2017_Structure_25_697 |
| PubMedID: 28392259 |
| Gene_locus related to this paper: pseae-PA2934 |