NOAA
Product of hydrolyssi of Pseudomonas quinolone signal (PQS) by AqdC
General
Type : Benzoate
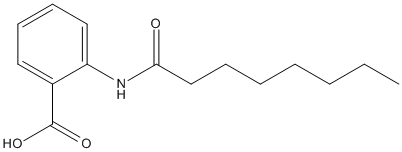
Chemical_Nomenclature : 2-(Octanoylamino)benzoic acid
Canonical SMILES : CCCCCCCC(=O)NC1=CC=CC=C1C(=O)O
InChI : InChI=1S\/C15H21NO3\/c1-2-3-4-5-6-11-14(17)16-13-10-8-7-9-12(13)15(18)19\/h7-10H,2-6,11H2,1H3,(H,16,17)(H,18,19)
InChIKey : DIBJPKPIMHBNDD-UHFFFAOYSA-N
Other name(s) : N-octanoyl anthranilic acid,n-octanoylanthranilic acid,AC1N4L6Y,N-n-octanoylanthranilic acid,AO-548\/41128890,SCHEMBL2183707,DIBJPKPIMHBNDD-UHFFFAOYSA-N,ZINC3157900,MCULE-7219401519
MW : 263.33
Formula : C15H21NO3
CAS_number :
PubChem : 4157220
UniChem : DIBJPKPIMHBNDD-UHFFFAOYSA-N
IUPHAR :
Wikipedia :
Target
Families : NOAA ligand of proteins in family: HOD-cofactorfree-dioxygenase
Stucture : 6RA3 Structural basis for recognition and ring-cleavage of the Pseudomonas quinolone signal (PQS) by AqDC in complex with its product
Protein : mycab-x8en65
References (2)
| Title : Structural basis for recognition and ring-cleavage of the Pseudomonas quinolone signal (PQS) by AqdC, a mycobacterial dioxygenase of the alpha\/beta-hydrolase fold family - Wullich_2019_J.Struct.Biol_207_287 |
| Author(s) : Wullich SC , Kobus S , Wienhold M , Hennecke U , Smits SHJ , Fetzner S |
| Ref : J Struct Biol , 207 :287 , 2019 |
| Abstract : Wullich_2019_J.Struct.Biol_207_287 |
| ESTHER : Wullich_2019_J.Struct.Biol_207_287 |
| PubMedSearch : Wullich_2019_J.Struct.Biol_207_287 |
| PubMedID: 31228546 |
| Gene_locus related to this paper: mycab-x8en65 |
| Title : Mycobacterium abscessus subsp. abscessus Is Capable of Degrading Pseudomonas aeruginosa Quinolone Signals - Birmes_2017_Front.Microbiol_8_339 |
| Author(s) : Birmes FS , Wolf T , Kohl TA , Ruger K , Bange F , Kalinowski J , Fetzner S |
| Ref : Front Microbiol , 8 :339 , 2017 |
| Abstract : Birmes_2017_Front.Microbiol_8_339 |
| ESTHER : Birmes_2017_Front.Microbiol_8_339 |
| PubMedSearch : Birmes_2017_Front.Microbiol_8_339 |
| PubMedID: 28303132 |
| Gene_locus related to this paper: rhoer-aqdC1 , rhoer-aqdC2 |